Gerrit Großmann
Dr. rer. nat., M. Sc., Postdoc at DFKI. Saarbrücken and Kaiserslautern. Germany.
Hello World! My name is Gerrit Großmann, welcome to my personal academic webpage! I work at the research department on Data Science and its Applications (DSA) (formerly Neuro-mechanistic Modeling) at the German Research Center for Artificial Intelligence (DFKI). I am also a member of the Machine Learning & Global Health Network (MLGH).
Research Interests:
🤖💡 My research revolves around the question: How can we integrate the distinct realms of discrete structures such as graphs and networks with the continuous nature of dynamic evolution, diffusion, and learning?
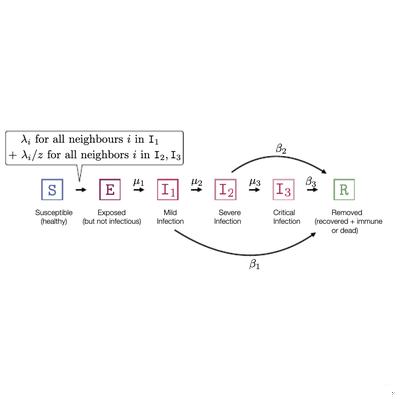
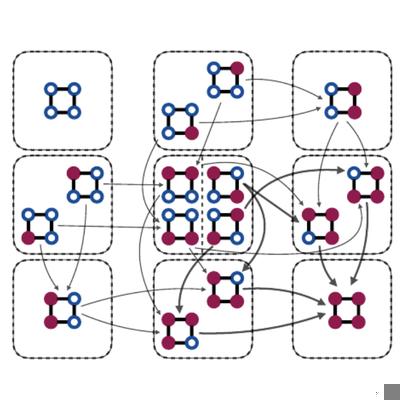
🎲🕸️ I am developing numerical methods to analyze spatio-temporal and stochastic dynamical processes. This research aims to understand how a substrate like a network shapes collective phenomena.
🧪🧠 Additionally, in collaboration with the NextAid project, my focus is on geometric deep learning for molecules. In this area, probabilistic flow models offer an innovative approach to integrating geometric deep learning with stochastic processes. My current projects include advancing neuro-symbolic guidance of diffusion models, implementing semi-supervised learning on metabolic networks, exploring the expressiveness of message-passing architectures, and the development of non-parametric methods for network reconstruction.
Please also check out our ★ STAR: Space × Time × Causality Reading Group.
Also, I am a mentor at ✦ AI Grid.
Overview:
Short CV
Teaching
- Engineering with Generative AI (WS 24/25) (Lecture at RPTU)
- Bridging the Gap - Language Models and Structured Knowledge in AI (SoSe 24)
- Advanced Topics in Diffusion Modeling: From Theory to Implementation (WS 23/23)
- AI for the Social Good (SoSe 23)
- Deep Generative Diffusion Models (WS 22/23)
- Deep Learning for Drug Discovery (SoSe 22)
- Concurrent Programming (SoSe 21)
- Exploring Graph Neural Networks (SoSe 21)
- Statistics Lab (SoSe 20)
- Exploring Complex Network Dynamics (WS 19/20)
- Probabilistic Models and Data Analysis (SoSe 19)
- Ringvorlesung: Perspektiven der Informatik (WS 18/19)
- Probabilistic Models of Concurrency (WS 18/19)
- Pattern and Speech Recognition
Tutorials, Talks, and Opinion Pieces
- Talk on Generative AI for Drug Discovery: Dreaming New (Useful) Molecules as part of the Society of Spanish Scientists in Germany (CERFA) 2024 Symposium: “Excuse me, there is Artificial Intelligence in my Soup” (find the slides here)
- Talk and tutorial on Language Models and Structured Knowledge in AI as part of the Inria-DFKI European Summer School
- Talk on Molecule Generation with Graph Neural Networks and Probabilistic Diffusion as part of the ScaDS.AI Summer School (find the slides here).
- Tutorial on (Neural) Spatio-Temporal Models as part of the Collaborative Intelligence lecture at RPTU.
- LLM4Science: Current Trends and Future Prospects of a New Paradigm
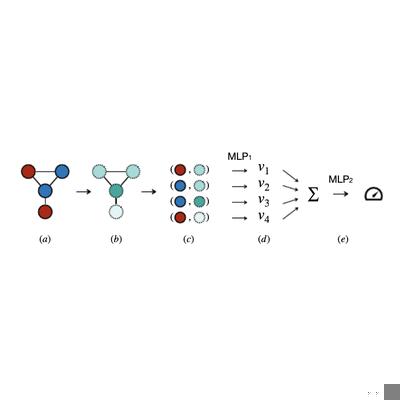
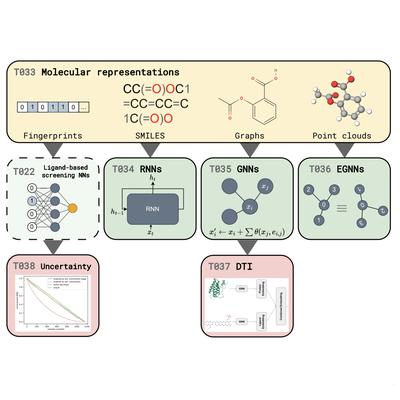
- I contributed to the deep learning for computer-aided drug design tutorial (T033 - T038) series.
- 7 Reasons Not to Use ODEs for Epidemic Modeling
- Der Nutzen von Corona-Modellen bleibt ungeklärt (The usefulness of corona models remains uncertain), Saarbrücker Zeitung; Feb., 4th, 2022
(Co-)Supervised Students
2026
Vaibhav Jain
Master's thesis
2026
Tanya Amit Tyagi
Master's thesis
2025
Daniel Schmalz
Bachelor's thesis
2025
Davronbek Islamov
Masters's thesis
2025
Zhifei Li
Masters's thesis
2025
Philipp Radder
Bachelor's thesis
2024
Prashanth Pombala
Master's thesis
2024
Anishka Singh
Bachelor's thesis
2024
Diya Narayanan
Bachelor's thesis
2024
Désirée Wiltzius
Bachelor's thesis
2024
Yehia Farghaly
Bachelor's thesis
2024
Magnus Cunow
Bachelor's thesis
2024
Nesara Belakere Lingarajaiah
Master's thesis
2023
Janine Lohse
Research Immersion Lab
2023
Joshgun Guliyev
Master's thesis
2022
Yan Yan Lau
Bachelor's thesis
2021
Julian Zimmerlin
Bachelor's thesis
2020
Lisa Heidmann
Bachelor's thesis
2020
Jonas Klesen
Research Immersion Lab
Theses
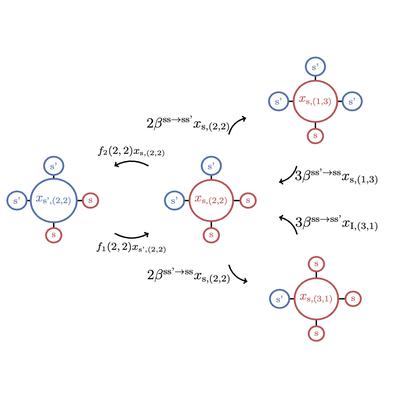
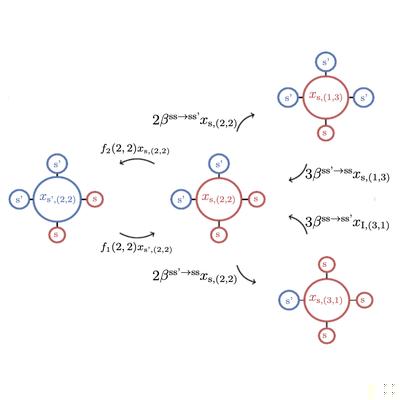
Lumping the Approximate Master Equation for Stochastic Processes on Complex Networks
Master’s Thesis
Avaliable upon request or at Campus-Bibliothek.
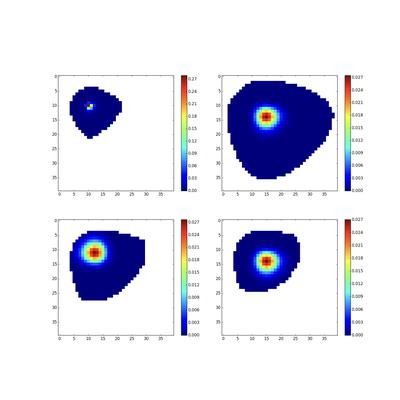
Efficient Computation of Likelihoods in Large Markov Models
Bachelor’s Thesis
Avaliable upon request or at Campus-Bibliothek.
Publications
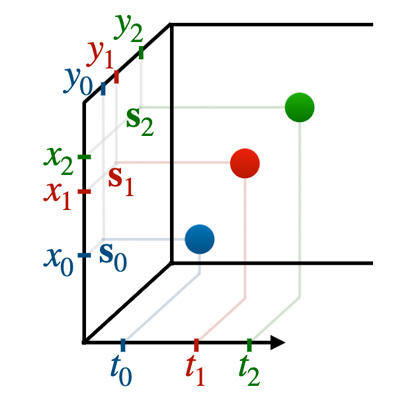
Neural Spatiotemporal Point Processes: Trends and Challenges
S. Mukherjee, M. Elhamdi, G. Mohler, D. Selby, Y. Xie, S. Vollmer, G. Großmann
Transactions on Machine Learning Research (TMLR), 2025

SQUID: A Bayesian Approach for Physics-Informed Event Modeling
S. Mukherjee, S. Vollmer, G. Großmann
Preprint, 2025
PDF ,
Accepted at EurIPS Differentiable Systems and Scientific Machine Learning Workshop, 2025
</div> </div> </div>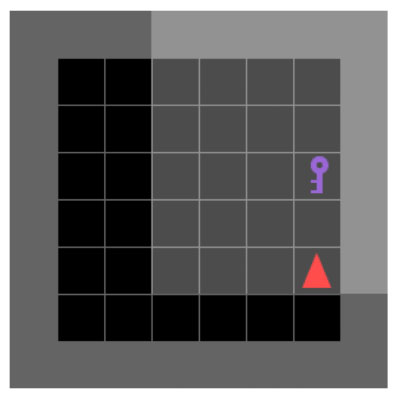
Guiding Exploration in Reinforcement Learning Through LLM-Augmented Observations
V. Jain, G. Großmann
Preprint, 2025
PDF , accepted at ICAPS LM4Plan Workshop
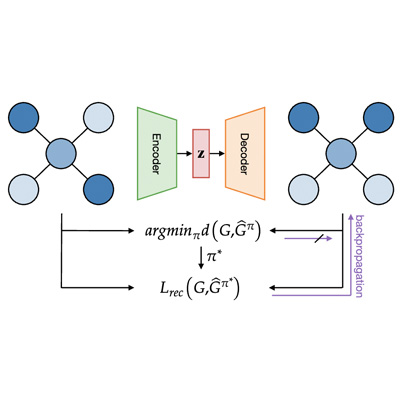
Auto-encoding Molecules: Graph-Matching Capabilities Matter
M. Cunow, G. Großmann, V. Wolf, S. Vollmer
Preprint, 2025
PDF , accepted at NeurIPS / ELLIS Generative Models, LLMs, and the Future of Molecular AI [ML4Molecules 2025] Workshop
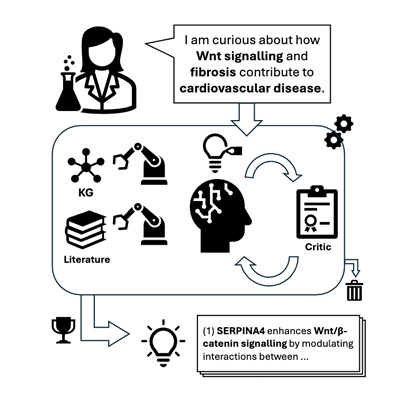
Biodisco: Multi-Agent Hypothesis Generation With Dual-Mode Evidence, Iterative Feedback and Temporal Evaluation
Y. Ke, K. George, K. Pandya, D. Blumenthal, M. Sprang, G. Großmann, S. Vollmer, D. Selby
Preprint, 2025

MEDAKA: Construction of Biomedical Knowledge Graphs Using Large Language Models
S. Sengupta, D. Selby, S. Vollmer, G. Großmann
Preprint, 2025
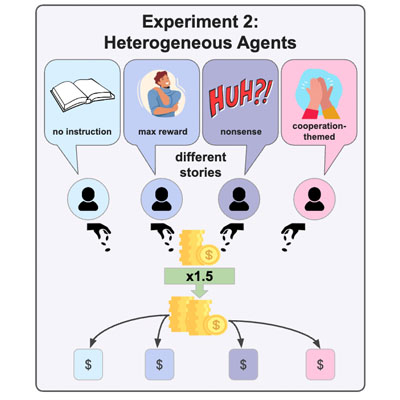
The Power of Stories: Narrative Priming Shapes How LLM Agents Collaborate and Compete
G. Großmann, L. Ivanova, S. L. Poduru, M. Tabrizian, I. Mesabah, D. Selby, S. Vollmer
NETGCOOP 2025, 2025
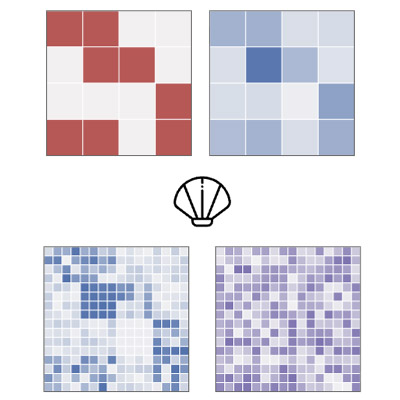
CLAM: Causal Spatial Disaggregation to Infer Local Effects From Coarse Data
G. Großmann, S. Mukherjee, S. Vollmer
Accepted at NeurIPS CauScien: Uncovering Causality in Science Workshop, 2025
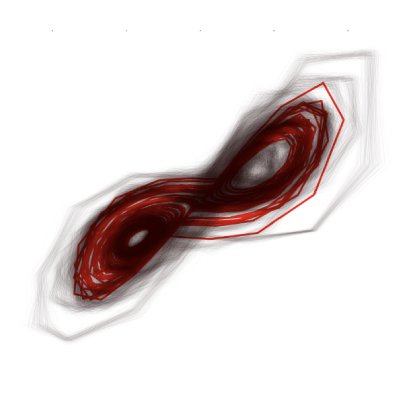
When Counterfactual Reasoning Fails: Chaos and Real-World Complexity
Y. Aalaila, G. Großmann, S. Mukherjee, J. Wahl, S. Vollmer
Preprint, 2025
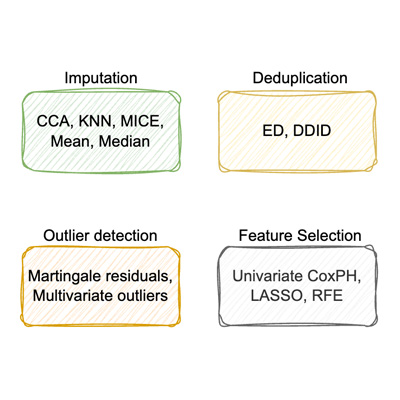
Cleansurvival: Automated Data Preprocessing for Time-To-Event Models Using Reinforcement Learning
Y. Koka, D. Selby, G. Großmann, S. Vollmer
Preprint, 2025
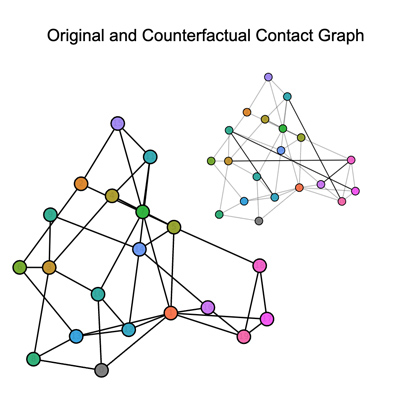
Peculiarities of Counterfactual Point Process Generation
G. Großmann, S. Mukherjee, S. Vollmer
Paper at STCausal, 2024
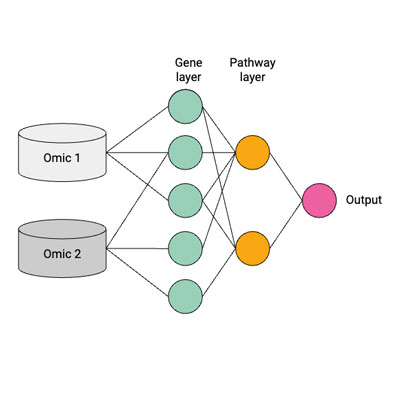
Visible Neural Networks for Multi-Omics Integration: A Critical Review
D. A. Selby, R. Jakhmola, M. Sprang, G. Großmann, H. Raki, N. Maani, D. Pavliuk, J. Ewald, S. Vollmer
Frontiers in Artificial Intelligence, 2024
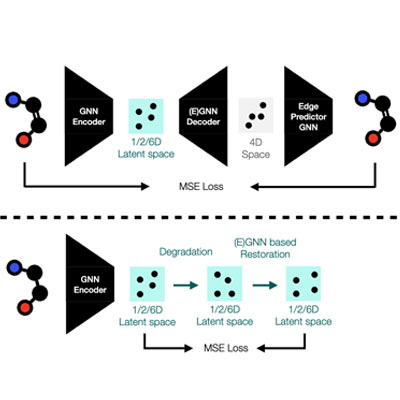
Exploring Molecule Generation Using Latent Space Graph Diffusion
P. Pombala, G. Grossmann, V. Wolf
Preprint, 2024
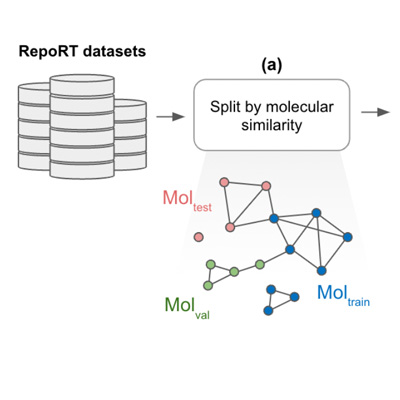
GRIP: Physics-Informed Neural Network for Gradient Retention Time Prediction in Liquid Chromatography
K. George, F.P. J. Haeckl, G. Großmann, A. Gurevich, A. Tagirdzhanov
Preprint, 2024
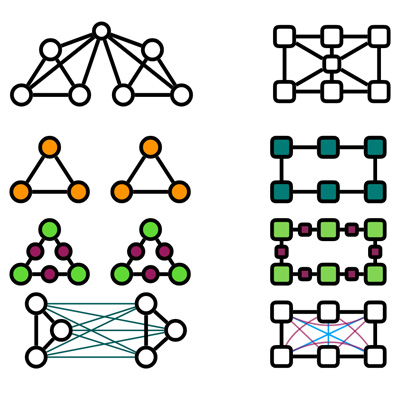
Enhancing GNNs with Architecture-Agnostic Graph Transformations: A Systematic Analysis
Z. Li, G. Großmann, V. Wolf
Paper at Complex Networks Conference, 2024
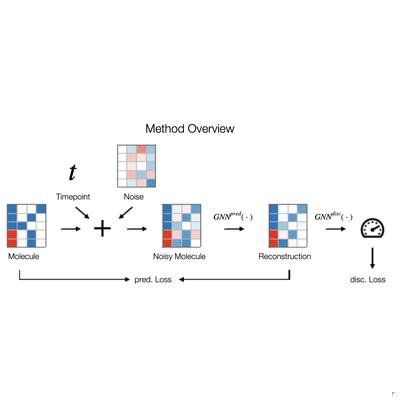
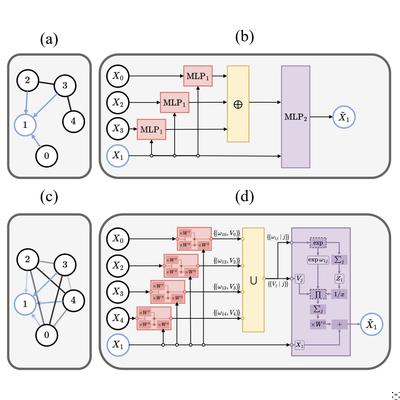
Elucidating the Relationship Between Transformers and GNNs
J. Groß, G. Großmann, V. Wolf
Preprint, 2023
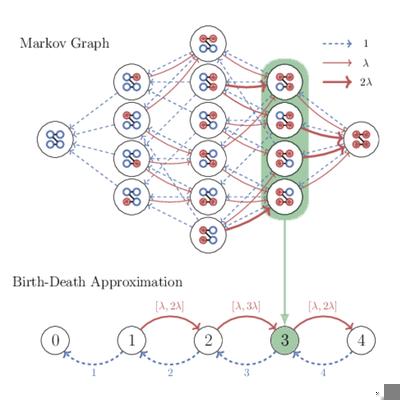
Birth-Death Processes Reproduce the Epidemic Footprint
G. Großmann, M. Backenköhler
Extended Abstract at Complex Networks Conference, 2022
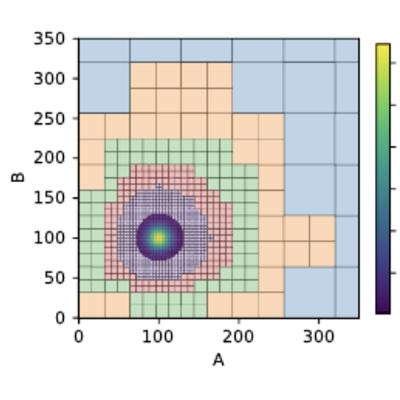
Abstraction-Guided Truncations for Stationary Distributions of Markov Population Models
M. Backenköhler, L. Bortolussi, G. Großmann, V. Wolf
Quantitative Evaluation of Systems Conference (QEST), 2021
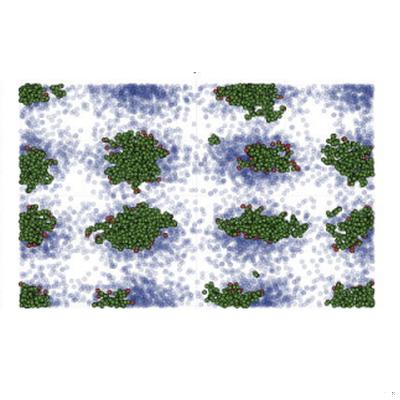

Epidemic Overdispersion Strengthens the Effectiveness of Mobility Restrictions
G. Großmann, M. Backenköhler, V. Wolf
Poster Abstract, 24th International Conference on Hybrid Systems: Computation and Control (HSCC), 2021
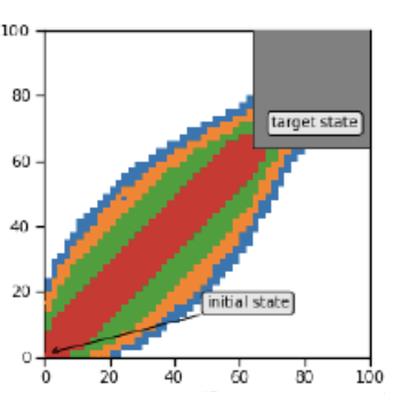
Analysis of Markov Jump Processes under Terminal Constraints
M. Backenköhler, L. Bortolussi, G. Großmann, V. Wolf
Tools and Algorithms for the Construction and Analysis of Systems Conference (TACAS), 2021
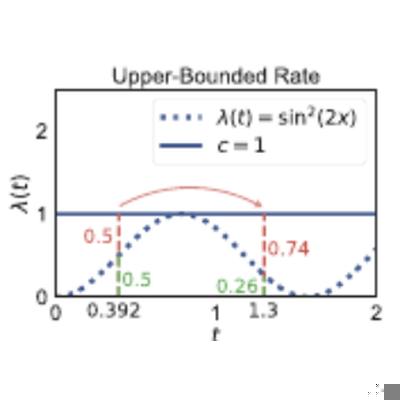
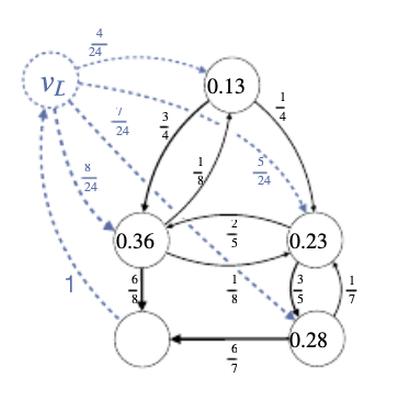
Learning Vaccine Allocation from Simulations
G. Großmann, M. Backenköhler, J. Klesen, V. Wolf
The 9th International Conference on Complex Networks and their Applications, 2020
Importance of Interaction Structure and Stochasticity for Epidemic Spreading: A COVID-19 Case Study
G. Großmann, M. Backenköhler, V. Wolf
Quantitative Evaluation of Systems Conference (QEST), 2020
Rejection-Based Simulation of Non-Markovian Agents on Complex Networks
G. Großmann, L. Bortolussi, V. Wolf
The 8th International Conference on Complex Networks and their Applications, 2019
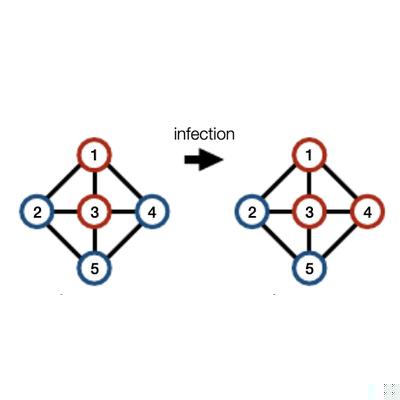
Rejection-Based Simulation of Stochastic Spreading Processes on Complex Networks
G. Großmann, V. Wolf
6th International Workshop on Hybrid Systems Biology (HSB), 2019
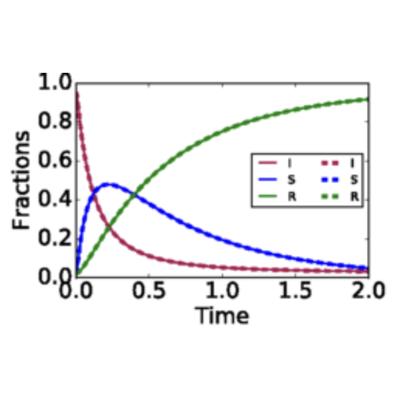
Lumping of Degree-Based Mean Field and Pair Approximation Equations for Multi-State Contact Processes
C. Kyriakopoulos, G. Großmann, V. Wolf, L. Bortolussi
PHYSICAL REVIEW E, 2019